Unsupervised Clustering for Nuclei Segmentation at Multiple Levels in H&E Histopathology Images
Overview
This project develops and evaluates three independent unsupervised methods for nuclei segmentation in H&E-stained histopathology images, a critical task for digital pathology. Each method leverages a different feature set—spatial, pixel-level, and stain-specific—to perform clustering and identify nuclei without the need for labeled data. By comparing these distinct approaches, the study identifies the most effective strategy for annotation-free nuclei detection. This work was awarded the 2nd Prize at the University of Hertfordshire's Data Science Project Club poster presentation for its rigorous comparative analysis and impactful results.
The Problem
In digital pathology, analyzing cell nuclei morphology is vital for disease grading, but supervised deep learning methods are constrained by their dependence on large, expert-annotated datasets. This annotation process is a significant bottleneck, limiting scalability and slowing down research. This project addresses this challenge by exploring and comparing different unsupervised clustering frameworks that can perform nuclei segmentation directly from raw image data, providing a scalable alternative to supervised techniques.
Approach & Technical Details
The core of this work was a comparative evaluation of three separate clustering-based methods using the MoNuSeg 2018 dataset. Each approach was designed to exploit different image features.
- Method 1: Superpixel-Level Clustering – Segments the image into superpixels (using SLIC) and applies clustering algorithms (K-Means, GMM, FCM) to identify coarse tissue regions, effective for high-level tissue mapping.
- Method 2: Pixel-Level Clustering – Operates directly on raw pixel data, clustering pixels based on color and intensity to refine localization of potential nuclear regions.
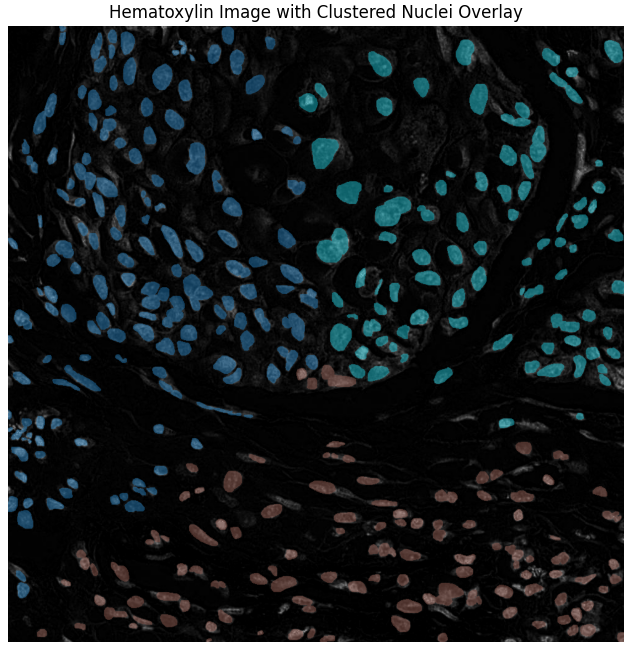
- Method 3: Stain-Specific Clustering (Hematoxylin Channel) – Uses color deconvolution (Macenko normalization) to separate the Hematoxylin stain, performing clustering solely on this channel for accurate nuclei detection.
Key Technologies: Python | Unsupervised Learning | Clustering (K-Means, GMM, FCM) | SLIC Superpixels | Color Deconvolution (Macenko Normalization) | Digital Histopathology (MoNuSeg Dataset)
Results & Outcomes
- Superior Performance of Stain-Specific Method: Method 3 yielded the most accurate and reliable instance-level nuclei segmentation.
- Effective Unsupervised Segmentation: Unsupervised clustering proved to be a viable alternative when labeled data is scarce.
- Award-Winning Research: Recognized with the 2nd Prize at the University of Hertfordshire Data Science Project Club poster presentation.
- Foundation for Scalable Pathology Tools: Offers insights for developing scalable digital pathology pipelines.